$1.85M NIH grant funds project to study virus interaction with the immune system and identify poxvirus threats
Wednesday, Sep. 9, 2015
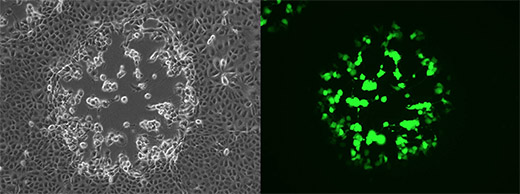
An African green monkey cell line was infected with a vaccinia virus strain that encodes a green fluorescent protein for the easy visualization of infection. Twenty-four hours after infection, plaque formation in the cell layer was observed, which resulted from cells that were destroyed by the virus, pictured in the left image. Virus-infected cells appear green in image obtained using fluorescence microscopy, pictured in the right image. Images captured by Sherry Haller in the Rothenburg lab. | Download the following image.
MANHATTAN — The National Institutes of Health are funding a Kansas State University research project that is looking at viruses that have the potential to be the next smallpox as well as an effective weapon against cancer.
Stefan Rothenburg, assistant professor of biology, was recently awarded more than $1.85 million in funding from the NIH for the project "Importance of Species-Specific Interactions of PKR with Poxvirus Inhibitors for Virus Replication and Host Range."
The project looks at the family of viruses classified as poxviruses and the biological factors that determine whether a poxvirus has the potential to infect different animal species and humans. A virus's interaction with the immune system determines whether a virus will be infectious to a species as well as how many different species the virus can infect, Rothenburg said.
Understanding how and why certain viruses replicate in certain hosts will help scientists to better predict possible future hosts and assess which viruses have strong potential for animal to human transmission. It also may lead to new strategies for disease control.
"Right now we have no way of knowing if a virus that evolves in an animal can infect another animal species and pose a serious threat to human health," Rothenburg said. "For example, influenza viruses naturally infect water fowl, but also can infect other species such as chickens, pigs and humans. Our current knowledge of how viruses interact with our immune system is insufficient to accurately predict if newly emerging influenza viruses pose serious human health threats."
A majority of emerging diseases that infect people began as animal viruses, Rothenburg said. In order to be prepared for those diseases, researchers have to study these viruses and determine their potential to become human pathogens.
One aspect of the project has Rothenburg and his lab trying to identify poxvirus threats.
"Smallpox, which was caused by the variola virus, was the most dangerous infectious disease ever known to mankind," Rothenburg said. "In human history, more people have died from smallpox infections than of all other infectious diseases combined, including AIDS, malaria and tuberculosis."
The ancestor of the variola virus, which causes smallpox, began as an animal disease and evolved into a highly infectious human virus. The mortality rate for smallpox ranged between 10-30 percent, and the virus spread for thousands of years and infected billions of people before the systematic use of a vaccine led to its eradication from nature. Blindness and disfiguring scars often afflicted survivors of smallpox.
Rothenburg is not conducting research with smallpox — a biosafety level-4 disease that only exists in two high security facilities in the world. Rather, experiments are being conducted with vaccinia virus, a weakened virus closely related to smallpox that served as the smallpox vaccine.
Rothenburg and researchers in his laboratory are taking genes from 16 different poxviruses, including the vaccinia and variola viruses, and inserting them into cells to observe how the viral proteins interact with the immune system of the host cells. They are looking at the behavior of PKR, a protein in cells that is activated when it detects a virus.
In previous studies, Rothenburg and other researchers found that PKR could stop the virus's replication and spread. Further research has found that although the PKR protein is found in the cells of all mammals, its sequence varies greatly even between closely related species, and the effectiveness of PKR in inhibiting certain viruses differs greatly depending on the species. For example, mice and rats are both rodents, but mouse PKR was found to be susceptible to certain poxvirus inhibitors whereas rat PKR was resistant.
"On a molecular level, we have no idea why vaccinia virus has such a broad range of hosts, but is not very virulent to humans while the smallpox virus infection was restricted to humans in which it is highly virulent," Rothenburg said.
Shedding light on the amino acids in PKR that give each species resistance or susceptibility to a particular poxvirus may help scientists predict which viruses pose a significant public health threat in the future.
"In the past two years a couple of new poxvirus that cause infections in humans have been identified," Rothenburg said. "Most poxviruses, though, are not considered big threats. We saw with smallpox that these viruses can evolve to infect humans and cause an epidemic. Evolution of another highly pathogenic poxvirus will happen sooner or later, so to be prepared we have to study these viruses and seek out what potential they have to become human pathogens."
Additionally, understanding this molecular machinery may lead to new advances in cancer research. Poxviruses, for example, have been shown to infect and kill certain types of human cancer cells. The research from the Rothenburg lab will help determine how poxviruses can be modified to improve their capabilities to selectively kill human cancer cells.
Microbiology doctoral students Sherry Haller, Topeka, and Chen Peng, China, conducted research that contributed to the preliminary data for the project. Kansas State University's Johnson Cancer Research Center partly funded the experiments, which Rothenburg said led to the awarded NIH grant.
The Division of Biology is in the College of Arts & Sciences.
